FDA-GPT
Description
FDA-GPT is a semantic search engine designed to perform document retrieval and Question Answering (QA) on a collection of FDA approval packets. Essentially, it helps users find relevant information and answers from a large set of FDA documents.
This wiki will focus on the abstract design of the application, making it easier for the infrastructure team to support and expand it.
High-Level Design
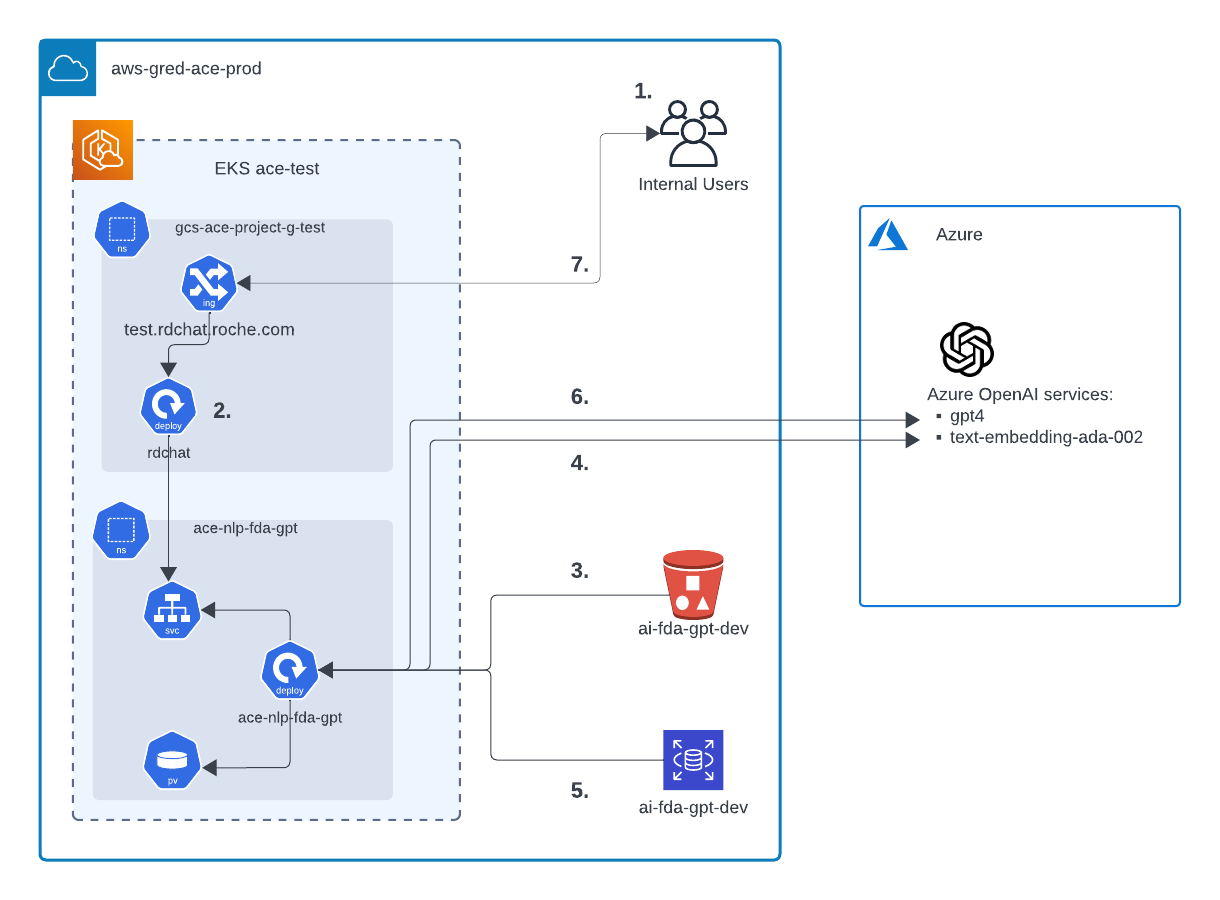 src: Lucidchart
src: Lucidchart
-
User Access: Users access FDA-GPT through test.rdchat.roche.com by selecting FDA-GPT as the model.
-
Connection to fda-chat Pod:
rdchatconnects to thefda-chatpod viaClusterIP. -
Metadata Extraction Model: The Metadata Extraction Model uses a checkpoint to load a pre-trained Named Entity Recognition (NER) model. This model extracts drug names from user queries to filter relevant documents. The checkpoint for the Metadata Extraction Model can be found at:
ai-fda-gpt-dev-bucket/ddi_drug_model.zip. This zip file is downloaded by aninitContainerto a Persistent Volume (PV) when the pod starts and is mounted under/model_artifacts/ddi_drug_model. -
Embedding Generation: The
text-embedding-ada-002model generates embeddings for user queries. -
Vector Storage and Search: Vector embeddings for FDA document chunks are stored in a PostgreSQL database with the pgvector extension. The system enables searching through the database using SQL queries to compare the user query vector to the FDA document vectors.
-
Conversational QA: The
gpt-4model handles the conversational QA component, summarizing the documents retrieved based on the user query. -
Response: The summarized response is sent back to the user.
The FDA pod is running on GPU Nodes.
Team Responsible
The team responsible for maintaining and troubleshooting the application.
Main Point of Contact:
- Jaymin Soni
- Email: JAYMIN.SONI@CONTRACTORS.ROCHE.COM
- unix-id:
sonij3
- Data Engineering Team
- Slack:
#ecdi-ace-data-engineering - Slack:
#fda-gpt-ace
- Slack:
GitHub
The repository for the application can be found at github.com/gred-ecdi/ace-nlp-fda-gpt.
Deployment Information
| Staging | Production | |
|---|---|---|
| git branch | main | - |
| Deployment environment | ace-test (EKS) Namespace: ace-nlp-fda-gpt | - |
| Deployment tool | https://argocd.eks.test.gred.ai/applications/argocd/ace-nlp-fda-gpt | - |
| URL | via test.rdchat.roche.com | - |
| S3 Bucket | - ai-fda-gpt-dev-bucket - Application connects to s3 via ai-fda-gpt-dev-read-app-fda-gpt s3 access point | - |
| Service Account | - serviceAccount Name: ace-nlp-fda-gpt - irsa role: arn:aws:iam::712649426017:role/irsa.ace-test.ace-nlp-fda-gpt.fda | - |
The Terraform workspace for the infrastructure can be found at terraform-ace-prod/us-west-2/ai-fda-gpt-dev.
How does the application get access to its s3 bucket ?
EKS Service account -> IRSA IAM Role -> The role has the tag id:app -> IAM Policy ace-eks-app-s3-access
Branching Flow
How does Main Branch get versions?
If there is a ‘fix’, ‘feat’, or ‘hotfix’ present in the git commit, the GitHub action release will bump up the version. So it’s automatically.
How does helm Chart get new versions?
You should manually update the helm verion in Chart.yaml.
Monitoring
-
Monitoring Dashboard: Grafana
-
Metrics being monitored: Memory usage by pod, CPU usage by pod, Replica Availability, PV Capacity
-
Alerts and notifications:
-
-
Logging Dashboard: Logs are available on the same Grafana Dashboard.
- To search logs in more details, please log in to OpenSearch > EKS ACE Test - Applications Dashboard > then choose namespace
ace-nlp-fda-gpt
- To search logs in more details, please log in to OpenSearch > EKS ACE Test - Applications Dashboard > then choose namespace
TODOs
- Database data is currently saved via the master admin. Create a dedicated app user for this purpose.
- The
text-embedding-ada-002model is outdated. Update to the latest version. - Improve the CI/CD pipeline.